SAP 定义:
Ref.1: Murray, T. A 等人 2016 年发表的文章的解释:
Stretch-attend posture (SAP) occurs during risk assessment and is prevalent in common rodentce behavioral tests.
– Holly, K. S., Orndorff, C. O., & Murray, T. A. (2016). MATSAP: An automated analysis of stretch-attend posture in rodent behavioral experiments. Scientific reports , 6 (1), 31286.
MATSAP: An automated analysis of stretch-attend posture in rodent behavioral experiments
Ref.2: mousebehavior.org 网站的解释:
Stretched attend is an investigatory posture that is typically seen when the mouse is investigating a potentially dangerous stimulus or environment. The posture serves to maintain the body in a safe location, and to allow rapid retreat, while extending the head to investigate.
根据上述释义,当动物在探索潜在危险的刺激或环境时,通常会看到这种姿势。这个姿势是为了保持身体在一个安全的位置,并允许快速撤退,同时延伸头部进行探索。
上述 Ref.1 提供了一个工具包 MATSAP 用于计算 SAP,不过使用起来相对麻烦。如果基于BehaviorAtlas的 3D 数据如何计算 SAP 呢?下面给出详细教程。
数据准备
-
首先,如果跑完了 BehaviorAtlas Analyzer,你会在工程里面得到 【results\BeAOutputs】文件夹,里面存放着每个样本的 .h5 格式的结果文件。
-
h5 文件中存放着 3D 的身体坐标数据
代码主脚本
clc; clear; close all;
% Set working path -> [results\BeAOutputs]
current_path = 'Y:\huangkang\data_processing\demo\';
% You can modify this threshold based on experience and actual situation
ecc_thr = 0.89; % default 0.89.
speed_thr = 80; % default 80 mm/s.
fps = 30; % video fps
show_ellipse = false;
show_SAP = true;
% Load body trajectory for all samples
data_3dskls = load_3dskl_h5([current_path, 'results\BeAOutputs\']);
n_sample = size(data_3dskls, 1);
% Process each sample
for ism = 1:n_sample
body_parts_3d = data_3dskls.data3d{ism};
SAP_flag = cal_SAP_3d(body_parts_3d, show_ellipse, ecc_thr, speed_thr, fps, show_SAP);
pause
end
load_3dskl_h5 函数批量加载 3D 身体坐标数据
function data_3dskls = load_3dskl_h5(skl_path)
skl_files = dir([skl_path,'*results.h5']);
data_name = natsort({skl_files.name});
data_name = data_name';
n_data = length(data_name);
data_3dskls = struct();
back_idx = 13;
for id = 1:n_data
data = h5read([skl_path, cell2mat(data_name(id))], '/3Dskeleton/data3D')';
body_part_name = deblank(h5read([skl_path, cell2mat(data_name(id))], '/3Dskeleton/Bodyparts'))';
fps = double(h5read([skl_path, cell2mat(data_name(id))], '/3Dskeleton/FPS'));
offset_X = (max(data(:,back_idx*3-2)) + min(data(:,back_idx*3-2)))/2;
offset_Y = (max(data(:,back_idx*3-1)) + min(data(:,back_idx*3-1)))/2;
sample_name = data_name{id};
sample_name = split(sample_name, '_');
sample_name = sample_name{1};
data_3dskls(id).sample_name = sample_name;
group_label = split(sample_name, '-');
group_label = group_label{3};
data_3dskls(id).group_label = group_label;
data_3dskls(id).data3d = data;
posX = 500*normalize(data(:,1:3:end), 'range');
posY = 500*normalize(data(:,2:3:end), 'range');
data_3dskls(id).x = posX;
data_3dskls(id).y = posY;
data_3dskls(id).z = data(:,3:3:end);
data_3dskls(id).fps = fps;
data_3dskls(id).body_part_name = {body_part_name};
disp(['load_3dskl: ', num2str(id)]);
% hold on
% scatter(data_3dskls(id).x(:,13),data_3dskls(id).y(:,13))
%
% drawnow
end
data_3dskls = struct2table(data_3dskls);
cal_SAP_3d 计算 SAP 函数
function SAP_flag = cal_SAP_3d(body_parts_3d, show_ellipse, ecc_thr, speed_thr, fps, show_SAP)
% body_parts_3d: 3D coordinates of mouse body parts, N*M array, N is the
% numer of frame, M/3 is the number of body parts.
% SAP_flag: N*1 boolean array
n_frame = size(body_parts_3d, 1);
eccentricity = zeros(n_frame, 1);
back_idx = 13; % back point index
if show_ellipse
figure
end
disp('Calculating eccentricity, please wait!')
for ifi = 1:n_frame
% Calculate eccentricity
bp_x = body_parts_3d(ifi, 1:3:42);
bp_y = body_parts_3d(ifi, 2:3:42);
if show_ellipse
h1 = subplot(121);
mesh_mouse(body_parts_3d, ifi);
axis equal; view([0, 90]); zlim([-50, 50]);
hold on;
title('Ellipse fitting for 3D mouse (Top view)')
h2 = subplot(122);
mesh_mouse(body_parts_3d, ifi);
axis equal; view([-50, 30]); zlim([-50, 50]);
hold on;
title('Ellipse fitting for 3D mouse (3D view)')
end
if show_ellipse
subplot(121)
fit_ellipse(bp_x, bp_y, h1);
subplot(122)
ellipse_t = fit_ellipse(bp_x, bp_y, h2);
pause(0.01);
cla(h1); cla(h2);
else
ellipse_t = fit_ellipse(bp_x, bp_y);
end
if ~isempty(ellipse_t.a)
a = ellipse_t.long_axis;
b = ellipse_t.short_axis;
if a == 0
eccentricity(ifi) = 0;
else
eccentricity(ifi) = sqrt(a^2 - b^2)/a;
end
else
eccentricity(ifi) = eccentricity(ifi - 1);
end
end
eccentricity_sm = smooth(eccentricity, fps*1/2);
% Calculate speed
body_3d = fillmissing(body_parts_3d, 'linear'); % mat 格式
X = body_3d(:, back_idx*3-2);
Y = body_3d(:, back_idx*3-1);
mean_pos = [X, Y];
speed_raw = diff(mean_pos);
% 2D speed
speed2 = sqrt(speed_raw(:, 1).^2 + speed_raw(:, 2).^2);
speed2 = speed2/(1/fps);
speed2 = [speed2(1); speed2];
speed_sm2 = smooth(speed2, fps*1/2);
SAP_flag = speed_sm2 < speed_thr & eccentricity_sm > ecc_thr;
if show_SAP
ts = linspace(1/fps, n_frame/fps, n_frame);
figure
subplot(311)
plot(ts, speed_sm2)
xlabel('Time (s)'); ylabel('Speed (mm/s)')
subplot(312)
plot(ts, eccentricity_sm)
xlabel('Time (s)'); ylabel('Eccentricity')
subplot(313)
plot(ts, SAP_flag)
xlabel('Time (s)'); ylabel('SAP')
end
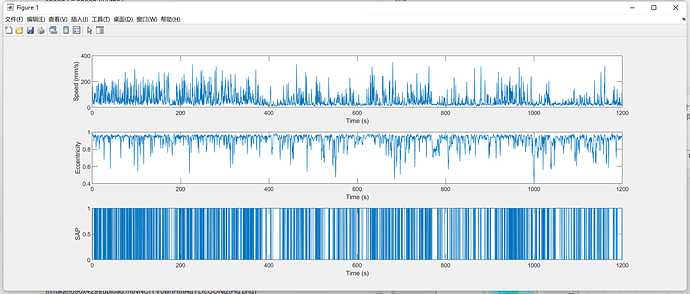
这个函数几个关键步骤,一是根据小鼠 X-Y 平面做椭圆拟合,计算 eccentricity 偏心率;然后计算小鼠的 X-Y 平面的位移速度。当某一帧 speed_sm2 < speed_thr & eccentricity_sm > ecc_thr 同时满足时,即可判定为 SAP。
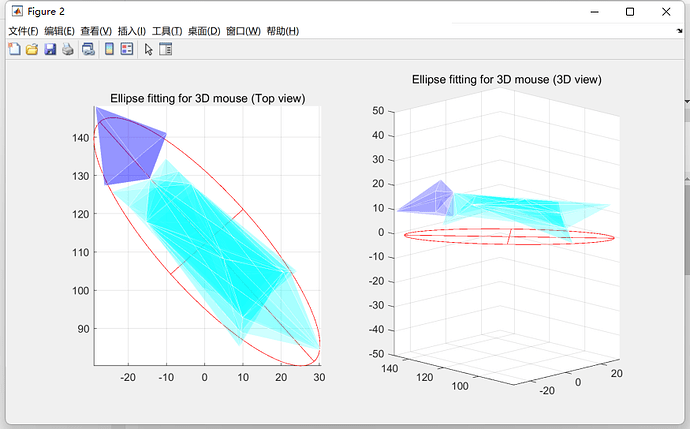
3D 小鼠椭圆拟合的效果:
SAP 的计算结果: